Ev ve Ofis taşıma sektöründe lider olmak.Teknolojiyi klrd takip ederek bunu müşteri menuniyeti amacı için kullanmak.Sektörde marka olmak.
İstanbul evden eve nakliyat
Misyonumuz sayesinde edindiğimiz müşteri memnuniyeti ve güven ile müşterilerimizin bizi tavsiye etmelerini sağlamak.
Video-based kinetic analysis of calcification in live osteogenic human embryonic stem cell cultures reveals the developmentally toxic effect of Snus tobacco extract
.png) Epidemiological studies suggest tobacco consumption as a probable environmental factor for a variety of congenital anomalies, including low bone mass and increased fracture risk. Despite intensive public health initiatives to publicize the detrimental effects of tobacco use during pregnancy, approximately 10–20% of women in the United States still consume tobacco during pregnancy, some opting for so-called harm-reduction tobacco. These include Snus, a type of orally consumed yet spit-free chewing tobacco, which is purported to expose users to fewer harmful chemicals. Concerns remain from a developmental health perspective since Snus has not reduced overall health risk to consumers and virtually nothing is known about whether skeletal problems from intrauterine exposure arise in the embryo.
Epidemiological studies suggest tobacco consumption as a probable environmental factor for a variety of congenital anomalies, including low bone mass and increased fracture risk. Despite intensive public health initiatives to publicize the detrimental effects of tobacco use during pregnancy, approximately 10–20% of women in the United States still consume tobacco during pregnancy, some opting for so-called harm-reduction tobacco. These include Snus, a type of orally consumed yet spit-free chewing tobacco, which is purported to expose users to fewer harmful chemicals. Concerns remain from a developmental health perspective since Snus has not reduced overall health risk to consumers and virtually nothing is known about whether skeletal problems from intrauterine exposure arise in the embryo.
Multi-label classification of stem cell microscopy images using deep learning
.png) This paper develops a pattern recognition and machine learning system to localize cell colony subtypes in multi-label, phase-contrast microscopy images. A convolutional neural network is trained to recognize homogeneous cell colonies, and is used in a sliding-window patch based testing method to localize these homogeneous cell types within heterogeneous, multi-label images. The method is used to determine the effects of nicotine on induced pluripotent stem cells expressing the Huntington’s disease phenotype. The results of the network are compared to those of an ECOC classifier trained on texture features.
This paper develops a pattern recognition and machine learning system to localize cell colony subtypes in multi-label, phase-contrast microscopy images. A convolutional neural network is trained to recognize homogeneous cell colonies, and is used in a sliding-window patch based testing method to localize these homogeneous cell types within heterogeneous, multi-label images. The method is used to determine the effects of nicotine on induced pluripotent stem cells expressing the Huntington’s disease phenotype. The results of the network are compared to those of an ECOC classifier trained on texture features.
DeephESC: An automated system for generating and classification of human embryonic stem cells
.png) In this paper, we introduce a hierarchical classification system consisting of Convolutional Neural Networks (CNN) and Triplet CNN’s to classify hESC images into six different classes. We also design an ensemble of Generative Adversarial Networks (GAN) for generating synthetic images of hESC’s. We validate the quality of the generated hESC images by training all of our CNN’s exclusively on the synthetic images generated by the GAN’s and evaluating them on the original hESC images.
In this paper, we introduce a hierarchical classification system consisting of Convolutional Neural Networks (CNN) and Triplet CNN’s to classify hESC images into six different classes. We also design an ensemble of Generative Adversarial Networks (GAN) for generating synthetic images of hESC’s. We validate the quality of the generated hESC images by training all of our CNN’s exclusively on the synthetic images generated by the GAN’s and evaluating them on the original hESC images.
3D Reconstruction of phase contrast images using focus measures
.png) In this paper, we present an approach for 3D phase-contrast microscopy using focus measure features. By using fluorescence data from the same location as the phase contrast data, we can train supervised regression algorithms to compute a depth map indicating the height of objects in imaged volume. From these depth maps, a 3D reconstruction of phase contrast images can be generated. This paper has shown the ability to reconstruct 3D phase contrast images using a variance metric inspired by all-in-focus methods. The proposed method has been used on A549 lung epithelial cells.
In this paper, we present an approach for 3D phase-contrast microscopy using focus measure features. By using fluorescence data from the same location as the phase contrast data, we can train supervised regression algorithms to compute a depth map indicating the height of objects in imaged volume. From these depth maps, a 3D reconstruction of phase contrast images can be generated. This paper has shown the ability to reconstruct 3D phase contrast images using a variance metric inspired by all-in-focus methods. The proposed method has been used on A549 lung epithelial cells.
HESCNET: A synthetically pre-trained convolutional neural network for human embryonic stem cell colony classification
.png) This paper proposes a method for improving the results of deep convolutional neural network classification using synthetic image samples. Generative adversarial networks are used to generate synthetic images from a dataset of phase contrast, human embryonic stem cell (hESC) microscopy images. hESCnet, a deep convolutional neural network is trained, and the results are shown on various combinations of synthetic and real images in order to improve the classification results with minimal data.
This paper proposes a method for improving the results of deep convolutional neural network classification using synthetic image samples. Generative adversarial networks are used to generate synthetic images from a dataset of phase contrast, human embryonic stem cell (hESC) microscopy images. hESCnet, a deep convolutional neural network is trained, and the results are shown on various combinations of synthetic and real images in order to improve the classification results with minimal data.
Extraction of Blebs in Human Embryonic Stem Cell Videos
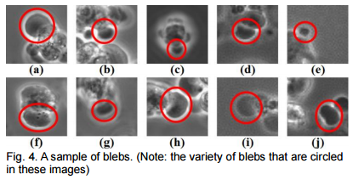 Analyzed are various segmentation methods for bleb extraction in hESC videos which introduces a bio-inspired score function to improve the performance in bleb extraction.
Full bleb formation consists of bleb expansion and retraction. Blebs change their size and image properties dynamically in both processes and between frames.
Therefore, adaptive parameters are needed for each segmentation method. A score function derived from the change of bleb area and orientation between consecutive frames is
proposed which provides adaptive parameters for bleb extraction in videos.
Analyzed are various segmentation methods for bleb extraction in hESC videos which introduces a bio-inspired score function to improve the performance in bleb extraction.
Full bleb formation consists of bleb expansion and retraction. Blebs change their size and image properties dynamically in both processes and between frames.
Therefore, adaptive parameters are needed for each segmentation method. A score function derived from the change of bleb area and orientation between consecutive frames is
proposed which provides adaptive parameters for bleb extraction in videos.
Evaluating cell processes, quality, and biomarkers in pluripotent stem cells using video bioinformatics
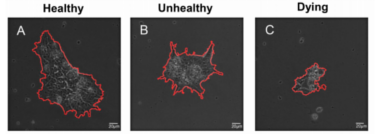 Presented is an unbiased and automated high-content profiling toolkit, StemCellQC, which non-invasively extracts information on
cell quality and cellular processes from time-lapse phase-contrast videos. Twenty four morphological and dynamic features were analyzed
in healthy, unhealthy, and dying human embryonic stem cell (hESC) colonies to identify those features that were affected in each group.
StemCellQC distinguished healthy and unhealthy/dying hESC colonies with 96% accuracy by non-invasively measuring and tracking dynamic
and morphological features over 48 hours. Changes in cellular processes were monitored by StemCellQC and predictions could be made
about the quality of pluripotent stem cell colonies. This toolkit reduced the time and resources required to track multiple pluripotent
stem cell colonies and eliminated handling errors and false classifications due to human bias.
Presented is an unbiased and automated high-content profiling toolkit, StemCellQC, which non-invasively extracts information on
cell quality and cellular processes from time-lapse phase-contrast videos. Twenty four morphological and dynamic features were analyzed
in healthy, unhealthy, and dying human embryonic stem cell (hESC) colonies to identify those features that were affected in each group.
StemCellQC distinguished healthy and unhealthy/dying hESC colonies with 96% accuracy by non-invasively measuring and tracking dynamic
and morphological features over 48 hours. Changes in cellular processes were monitored by StemCellQC and predictions could be made
about the quality of pluripotent stem cell colonies. This toolkit reduced the time and resources required to track multiple pluripotent
stem cell colonies and eliminated handling errors and false classifications due to human bias.
Bio-Driven Cell Region Detection in Human Embryonic Stem Cell Assay
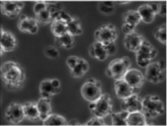 We present a bio-driven algorithm that detects cell regions automatically in the human
embryonic stem cell (hESC) images obtained using a phase contrast microscope. The intensity
distributions of foreground/hESCs and background/substrate are modelled as a mixture of two Gaussians.
In comparison with the state-of-the-art methods, the proposed method is able to detect the entire cell
region instead of fragmented cell regions. It also yields high marks on measures such as Jaccard similarity,
Dice coefficient, sensitivity and specificity.
We present a bio-driven algorithm that detects cell regions automatically in the human
embryonic stem cell (hESC) images obtained using a phase contrast microscope. The intensity
distributions of foreground/hESCs and background/substrate are modelled as a mixture of two Gaussians.
In comparison with the state-of-the-art methods, the proposed method is able to detect the entire cell
region instead of fragmented cell regions. It also yields high marks on measures such as Jaccard similarity,
Dice coefficient, sensitivity and specificity.
Comparison of Texture Features for Human Embryonic Stem Cells with Bio-Inspired Multi-Class Support Vector Machine
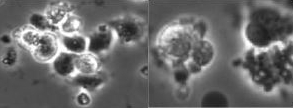 Determining the meaningful texture features for human embryonic stem cells (hESC) is important in the development of an online hESC classification
system. We propose the use of a novel support vector machine with bio-inspired one-against-all (OAA) multi-class structural and statistical
Gabor descriptors for hESC classification. We investigated the statistical histogram information at four different orientations and two different window
sizes of the Gabor filter. We've also demonstrated that statistical Gabor features are more accurate and reliable than conventional historgram based features.
Determining the meaningful texture features for human embryonic stem cells (hESC) is important in the development of an online hESC classification
system. We propose the use of a novel support vector machine with bio-inspired one-against-all (OAA) multi-class structural and statistical
Gabor descriptors for hESC classification. We investigated the statistical histogram information at four different orientations and two different window
sizes of the Gabor filter. We've also demonstrated that statistical Gabor features are more accurate and reliable than conventional historgram based features.
Automatic Cell Region Detection by K-means with Weighted Entropy
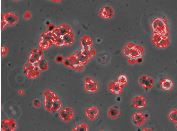 We propose an automatic method to detect human
embryonic stem cell regions. The proposed method
utilizes the K-means algorithm with weighted entropy. As
in phase contrast images the cell regions have high intensity
variation, they usually yield higher entropy values than the
substrate regions which have less intensity variation. Thus,
the entropy can be used as an important feature for the detection
of stem cells. However, homogeneity in intensity within
some of the cell bodies and halos surrounding the cell bodies
also gives low entropy values. Therefore, we introduce a
weighted entropy formulation which fuses entropy and image
intensity information to detect the entire cell regions.
We propose an automatic method to detect human
embryonic stem cell regions. The proposed method
utilizes the K-means algorithm with weighted entropy. As
in phase contrast images the cell regions have high intensity
variation, they usually yield higher entropy values than the
substrate regions which have less intensity variation. Thus,
the entropy can be used as an important feature for the detection
of stem cells. However, homogeneity in intensity within
some of the cell bodies and halos surrounding the cell bodies
also gives low entropy values. Therefore, we introduce a
weighted entropy formulation which fuses entropy and image
intensity information to detect the entire cell regions.
Automated Human Embryonic Stem Cell Detection
 We present an automated detection
method with simple algorithm for detecting human embryonic
stem cell (hESC) regions in phase contrast images. The
algorithm uses both the spatial information as well as the
intensity distribution for cell region detection. The method is
modeled as a mixture of two Gaussians; hESC and substrate
regions. The paper validates the method with various videos
acquired under different microscope objectives.
We present an automated detection
method with simple algorithm for detecting human embryonic
stem cell (hESC) regions in phase contrast images. The
algorithm uses both the spatial information as well as the
intensity distribution for cell region detection. The method is
modeled as a mixture of two Gaussians; hESC and substrate
regions. The paper validates the method with various videos
acquired under different microscope objectives.
Detection of Non-dynamic Blebbing Single Unattached Human Embryonic Stem Cells
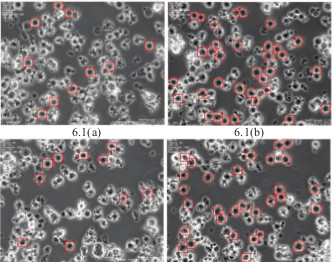 Human Embryonic Stem Cells (HESCs) are promising for
the treatment of many diseases and for toxicological testing.
There is a great interest among biologists to automatically
determine the number of various types of cells in a
population of mixed morphologies. This study addresses
quantification of non-dynamic blebbing single unattached
human embryonic stem cells (NDBSU-HESCs) that are in
suspension and do not show evidence of blebbing. Current
image processing methods are inadequate for detecting these
cells in real time. We propose a method for
NDBSU-HESC detection by using multiple trained
classifiers, where each classifier eliminates cells with
properties unmatched to NDBSU-HESCs. The paper
validates the method with many videos captured with live
stem cells.
Human Embryonic Stem Cells (HESCs) are promising for
the treatment of many diseases and for toxicological testing.
There is a great interest among biologists to automatically
determine the number of various types of cells in a
population of mixed morphologies. This study addresses
quantification of non-dynamic blebbing single unattached
human embryonic stem cells (NDBSU-HESCs) that are in
suspension and do not show evidence of blebbing. Current
image processing methods are inadequate for detecting these
cells in real time. We propose a method for
NDBSU-HESC detection by using multiple trained
classifiers, where each classifier eliminates cells with
properties unmatched to NDBSU-HESCs. The paper
validates the method with many videos captured with live
stem cells.
Human Embryonic Stem Cell Detection by Spatial Information and Mixture of Gaussians
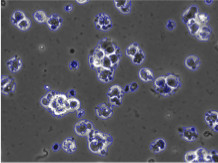 Human Embryonic Stem Cells (HESCs) possess
the potential to provide treatments for cancer, Parkinson's
disease, Huntington's disease, Type 1 diabetes, mellitus, etc. Consequently,
HESCs are often used in the biological assay to study
the effects of chemical agents in the human body. However,
detection of HESC is often a challenge in phase contrast images.
To improve the accuracy of HESC colony detection, we combine
spatial information and the outcome of a mixture of Gaussians
model. While a mixture of Gaussians generates reasonable
labels for various regions of HESC images, it lacks spatial
details and connectivity. Sets of spatially consistent candidate
labeling are generated by median filtering the image at different
scales followed by thresholding. An optimal combination of
filter scale and threshold which maximizes the correlation
coefficient between the spatial information and the mixture of
Gaussians output is obtained. The paper validates the method
for various HESC videos.
Human Embryonic Stem Cells (HESCs) possess
the potential to provide treatments for cancer, Parkinson's
disease, Huntington's disease, Type 1 diabetes, mellitus, etc. Consequently,
HESCs are often used in the biological assay to study
the effects of chemical agents in the human body. However,
detection of HESC is often a challenge in phase contrast images.
To improve the accuracy of HESC colony detection, we combine
spatial information and the outcome of a mixture of Gaussians
model. While a mixture of Gaussians generates reasonable
labels for various regions of HESC images, it lacks spatial
details and connectivity. Sets of spatially consistent candidate
labeling are generated by median filtering the image at different
scales followed by thresholding. An optimal combination of
filter scale and threshold which maximizes the correlation
coefficient between the spatial information and the mixture of
Gaussians output is obtained. The paper validates the method
for various HESC videos.
Video Bioinformatics Analysis of Human Embryonic Stem Cell Colony Growth
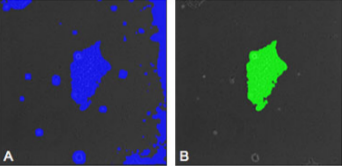 Mining information from video
material is difficult to do without the aid of computer software. In this article, we introduce a video
bioinformatics method for quantifying the growth of
human embryonic stem cells (hESC) by analyzing time-lapse videos. To determine the rate of growth
of these colonies,
three CL-Quant recipes were developed which enables users to extract various types of data
from video images. The first segmented
the image into the colony and background, the second enhanced the image to define colonies throughout the
video sequence accurately, and the third measured the number of pixels in the colony overtime.
When the data obtained using the CL-Quant
recipes and Photoshop were compared, results were virtually identical, indicating the CL-Quant recipes
were truthful. The method described here could be applied to any video data to measure growth rates of
hESC or other cells that grow in colonies.
Mining information from video
material is difficult to do without the aid of computer software. In this article, we introduce a video
bioinformatics method for quantifying the growth of
human embryonic stem cells (hESC) by analyzing time-lapse videos. To determine the rate of growth
of these colonies,
three CL-Quant recipes were developed which enables users to extract various types of data
from video images. The first segmented
the image into the colony and background, the second enhanced the image to define colonies throughout the
video sequence accurately, and the third measured the number of pixels in the colony overtime.
When the data obtained using the CL-Quant
recipes and Photoshop were compared, results were virtually identical, indicating the CL-Quant recipes
were truthful. The method described here could be applied to any video data to measure growth rates of
hESC or other cells that grow in colonies.
|


.png) Epidemiological studies suggest tobacco consumption as a probable environmental factor for a variety of congenital anomalies, including low bone mass and increased fracture risk. Despite intensive public health initiatives to publicize the detrimental effects of tobacco use during pregnancy, approximately 10–20% of women in the United States still consume tobacco during pregnancy, some opting for so-called harm-reduction tobacco. These include Snus, a type of orally consumed yet spit-free chewing tobacco, which is purported to expose users to fewer harmful chemicals. Concerns remain from a developmental health perspective since Snus has not reduced overall health risk to consumers and virtually nothing is known about whether skeletal problems from intrauterine exposure arise in the embryo.
Epidemiological studies suggest tobacco consumption as a probable environmental factor for a variety of congenital anomalies, including low bone mass and increased fracture risk. Despite intensive public health initiatives to publicize the detrimental effects of tobacco use during pregnancy, approximately 10–20% of women in the United States still consume tobacco during pregnancy, some opting for so-called harm-reduction tobacco. These include Snus, a type of orally consumed yet spit-free chewing tobacco, which is purported to expose users to fewer harmful chemicals. Concerns remain from a developmental health perspective since Snus has not reduced overall health risk to consumers and virtually nothing is known about whether skeletal problems from intrauterine exposure arise in the embryo.
.png) This paper develops a pattern recognition and machine learning system to localize cell colony subtypes in multi-label, phase-contrast microscopy images. A convolutional neural network is trained to recognize homogeneous cell colonies, and is used in a sliding-window patch based testing method to localize these homogeneous cell types within heterogeneous, multi-label images. The method is used to determine the effects of nicotine on induced pluripotent stem cells expressing the Huntington’s disease phenotype. The results of the network are compared to those of an ECOC classifier trained on texture features.
This paper develops a pattern recognition and machine learning system to localize cell colony subtypes in multi-label, phase-contrast microscopy images. A convolutional neural network is trained to recognize homogeneous cell colonies, and is used in a sliding-window patch based testing method to localize these homogeneous cell types within heterogeneous, multi-label images. The method is used to determine the effects of nicotine on induced pluripotent stem cells expressing the Huntington’s disease phenotype. The results of the network are compared to those of an ECOC classifier trained on texture features.
.png) In this paper, we introduce a hierarchical classification system consisting of Convolutional Neural Networks (CNN) and Triplet CNN’s to classify hESC images into six different classes. We also design an ensemble of Generative Adversarial Networks (GAN) for generating synthetic images of hESC’s. We validate the quality of the generated hESC images by training all of our CNN’s exclusively on the synthetic images generated by the GAN’s and evaluating them on the original hESC images.
In this paper, we introduce a hierarchical classification system consisting of Convolutional Neural Networks (CNN) and Triplet CNN’s to classify hESC images into six different classes. We also design an ensemble of Generative Adversarial Networks (GAN) for generating synthetic images of hESC’s. We validate the quality of the generated hESC images by training all of our CNN’s exclusively on the synthetic images generated by the GAN’s and evaluating them on the original hESC images.
.png) In this paper, we present an approach for 3D phase-contrast microscopy using focus measure features. By using fluorescence data from the same location as the phase contrast data, we can train supervised regression algorithms to compute a depth map indicating the height of objects in imaged volume. From these depth maps, a 3D reconstruction of phase contrast images can be generated. This paper has shown the ability to reconstruct 3D phase contrast images using a variance metric inspired by all-in-focus methods. The proposed method has been used on A549 lung epithelial cells.
In this paper, we present an approach for 3D phase-contrast microscopy using focus measure features. By using fluorescence data from the same location as the phase contrast data, we can train supervised regression algorithms to compute a depth map indicating the height of objects in imaged volume. From these depth maps, a 3D reconstruction of phase contrast images can be generated. This paper has shown the ability to reconstruct 3D phase contrast images using a variance metric inspired by all-in-focus methods. The proposed method has been used on A549 lung epithelial cells.
.png) This paper proposes a method for improving the results of deep convolutional neural network classification using synthetic image samples. Generative adversarial networks are used to generate synthetic images from a dataset of phase contrast, human embryonic stem cell (hESC) microscopy images. hESCnet, a deep convolutional neural network is trained, and the results are shown on various combinations of synthetic and real images in order to improve the classification results with minimal data.
This paper proposes a method for improving the results of deep convolutional neural network classification using synthetic image samples. Generative adversarial networks are used to generate synthetic images from a dataset of phase contrast, human embryonic stem cell (hESC) microscopy images. hESCnet, a deep convolutional neural network is trained, and the results are shown on various combinations of synthetic and real images in order to improve the classification results with minimal data.
 Presented is an unbiased and automated high-content profiling toolkit, StemCellQC, which non-invasively extracts information on
cell quality and cellular processes from time-lapse phase-contrast videos. Twenty four morphological and dynamic features were analyzed
in healthy, unhealthy, and dying human embryonic stem cell (hESC) colonies to identify those features that were affected in each group.
StemCellQC distinguished healthy and unhealthy/dying hESC colonies with 96% accuracy by non-invasively measuring and tracking dynamic
and morphological features over 48 hours. Changes in cellular processes were monitored by StemCellQC and predictions could be made
about the quality of pluripotent stem cell colonies. This toolkit reduced the time and resources required to track multiple pluripotent
stem cell colonies and eliminated handling errors and false classifications due to human bias.
Presented is an unbiased and automated high-content profiling toolkit, StemCellQC, which non-invasively extracts information on
cell quality and cellular processes from time-lapse phase-contrast videos. Twenty four morphological and dynamic features were analyzed
in healthy, unhealthy, and dying human embryonic stem cell (hESC) colonies to identify those features that were affected in each group.
StemCellQC distinguished healthy and unhealthy/dying hESC colonies with 96% accuracy by non-invasively measuring and tracking dynamic
and morphological features over 48 hours. Changes in cellular processes were monitored by StemCellQC and predictions could be made
about the quality of pluripotent stem cell colonies. This toolkit reduced the time and resources required to track multiple pluripotent
stem cell colonies and eliminated handling errors and false classifications due to human bias.
 We present a bio-driven algorithm that detects cell regions automatically in the human
embryonic stem cell (hESC) images obtained using a phase contrast microscope. The intensity
distributions of foreground/hESCs and background/substrate are modelled as a mixture of two Gaussians.
In comparison with the state-of-the-art methods, the proposed method is able to detect the entire cell
region instead of fragmented cell regions. It also yields high marks on measures such as Jaccard similarity,
Dice coefficient, sensitivity and specificity.
We present a bio-driven algorithm that detects cell regions automatically in the human
embryonic stem cell (hESC) images obtained using a phase contrast microscope. The intensity
distributions of foreground/hESCs and background/substrate are modelled as a mixture of two Gaussians.
In comparison with the state-of-the-art methods, the proposed method is able to detect the entire cell
region instead of fragmented cell regions. It also yields high marks on measures such as Jaccard similarity,
Dice coefficient, sensitivity and specificity.
 We present an automated detection
method with simple algorithm for detecting human embryonic
stem cell (hESC) regions in phase contrast images. The
algorithm uses both the spatial information as well as the
intensity distribution for cell region detection. The method is
modeled as a mixture of two Gaussians; hESC and substrate
regions. The paper validates the method with various videos
acquired under different microscope objectives.
We present an automated detection
method with simple algorithm for detecting human embryonic
stem cell (hESC) regions in phase contrast images. The
algorithm uses both the spatial information as well as the
intensity distribution for cell region detection. The method is
modeled as a mixture of two Gaussians; hESC and substrate
regions. The paper validates the method with various videos
acquired under different microscope objectives.
 Human Embryonic Stem Cells (HESCs) are promising for
the treatment of many diseases and for toxicological testing.
There is a great interest among biologists to automatically
determine the number of various types of cells in a
population of mixed morphologies. This study addresses
quantification of non-dynamic blebbing single unattached
human embryonic stem cells (NDBSU-HESCs) that are in
suspension and do not show evidence of blebbing. Current
image processing methods are inadequate for detecting these
cells in real time. We propose a method for
NDBSU-HESC detection by using multiple trained
classifiers, where each classifier eliminates cells with
properties unmatched to NDBSU-HESCs. The paper
validates the method with many videos captured with live
stem cells.
Human Embryonic Stem Cells (HESCs) are promising for
the treatment of many diseases and for toxicological testing.
There is a great interest among biologists to automatically
determine the number of various types of cells in a
population of mixed morphologies. This study addresses
quantification of non-dynamic blebbing single unattached
human embryonic stem cells (NDBSU-HESCs) that are in
suspension and do not show evidence of blebbing. Current
image processing methods are inadequate for detecting these
cells in real time. We propose a method for
NDBSU-HESC detection by using multiple trained
classifiers, where each classifier eliminates cells with
properties unmatched to NDBSU-HESCs. The paper
validates the method with many videos captured with live
stem cells.
 Human Embryonic Stem Cells (HESCs) possess
the potential to provide treatments for cancer, Parkinson's
disease, Huntington's disease, Type 1 diabetes, mellitus, etc. Consequently,
HESCs are often used in the biological assay to study
the effects of chemical agents in the human body. However,
detection of HESC is often a challenge in phase contrast images.
To improve the accuracy of HESC colony detection, we combine
spatial information and the outcome of a mixture of Gaussians
model. While a mixture of Gaussians generates reasonable
labels for various regions of HESC images, it lacks spatial
details and connectivity. Sets of spatially consistent candidate
labeling are generated by median filtering the image at different
scales followed by thresholding. An optimal combination of
filter scale and threshold which maximizes the correlation
coefficient between the spatial information and the mixture of
Gaussians output is obtained. The paper validates the method
for various HESC videos.
Human Embryonic Stem Cells (HESCs) possess
the potential to provide treatments for cancer, Parkinson's
disease, Huntington's disease, Type 1 diabetes, mellitus, etc. Consequently,
HESCs are often used in the biological assay to study
the effects of chemical agents in the human body. However,
detection of HESC is often a challenge in phase contrast images.
To improve the accuracy of HESC colony detection, we combine
spatial information and the outcome of a mixture of Gaussians
model. While a mixture of Gaussians generates reasonable
labels for various regions of HESC images, it lacks spatial
details and connectivity. Sets of spatially consistent candidate
labeling are generated by median filtering the image at different
scales followed by thresholding. An optimal combination of
filter scale and threshold which maximizes the correlation
coefficient between the spatial information and the mixture of
Gaussians output is obtained. The paper validates the method
for various HESC videos.
 Mining information from video
material is difficult to do without the aid of computer software. In this article, we introduce a video
bioinformatics method for quantifying the growth of
human embryonic stem cells (hESC) by analyzing time-lapse videos. To determine the rate of growth
of these colonies,
three CL-Quant recipes were developed which enables users to extract various types of data
from video images. The first segmented
the image into the colony and background, the second enhanced the image to define colonies throughout the
video sequence accurately, and the third measured the number of pixels in the colony overtime.
When the data obtained using the CL-Quant
recipes and Photoshop were compared, results were virtually identical, indicating the CL-Quant recipes
were truthful. The method described here could be applied to any video data to measure growth rates of
hESC or other cells that grow in colonies.
Mining information from video
material is difficult to do without the aid of computer software. In this article, we introduce a video
bioinformatics method for quantifying the growth of
human embryonic stem cells (hESC) by analyzing time-lapse videos. To determine the rate of growth
of these colonies,
three CL-Quant recipes were developed which enables users to extract various types of data
from video images. The first segmented
the image into the colony and background, the second enhanced the image to define colonies throughout the
video sequence accurately, and the third measured the number of pixels in the colony overtime.
When the data obtained using the CL-Quant
recipes and Photoshop were compared, results were virtually identical, indicating the CL-Quant recipes
were truthful. The method described here could be applied to any video data to measure growth rates of
hESC or other cells that grow in colonies.